from sklearn.compose import ColumnTransformer, make_column_selector
from sklearn.neighbors import KernelDensity
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import OneHotEncoder, PolynomialFeatures, SplineTransformer
from skcausal.causal_estimators import (
DirectNoCovariates,
DirectRegressor,
DoublyRobustPseudoOutcome,
GPS,
)
from skcausal.density.stabilized_from_conditional import (
KernelMarginalAndConditional,
)
def make_linear_pipeline():
preprocessor = ColumnTransformer(
transformers=[
(
"encode_categorical",
OneHotEncoder(
drop="first",
handle_unknown="ignore",
sparse_output=False,
),
make_column_selector(dtype_include=["category", "object", "string"]),
)
],
remainder="passthrough",
verbose_feature_names_out=False,
)
return make_pipeline(preprocessor, LinearRegression())
def make_gps_outcome_pipeline():
preprocessor = ColumnTransformer(
transformers=[
(
"expand_gps",
SplineTransformer(
n_knots=5,
degree=3,
include_bias=False,
),
["gps"],
),
(
"encode_categorical",
OneHotEncoder(
drop="first",
handle_unknown="ignore",
sparse_output=False,
),
make_column_selector(dtype_include=["category", "object", "string"]),
),
],
remainder="drop",
verbose_feature_names_out=False,
)
return make_pipeline(
preprocessor,
PolynomialFeatures(degree=2, include_bias=False),
LinearRegression(),
)
continuous_capable_results = {
"Truth": truth_curve,
"DirectNoCovariates": np.asarray(
DirectNoCovariates(outcome_regressor=make_linear_pipeline())
.fit(X, t, y)
.predict(t=levels),
dtype=float,
).reshape(-1),
"DirectRegressor": np.asarray(
DirectRegressor(outcome_regressor=make_linear_pipeline())
.fit(X, t, y)
.predict(t=levels),
dtype=float,
).reshape(-1),
"DoublyRobustPseudoOutcome": np.asarray(
DoublyRobustPseudoOutcome(
density_estimator=KernelMarginalAndConditional(
conditional_density_estimator=SklearnCategoricalDensity(
LogisticRegression(max_iter=1000)
),
kernel=KernelDensity(bandwidth=0.5),
),
outcome_regressor=make_linear_pipeline(),
pseudo_outcome_regressor=make_linear_pipeline(),
random_state=0,
)
.fit(X, t, y)
.predict(t=levels),
dtype=float,
).reshape(-1),
"GPS": np.asarray(
GPS(
density_regressor=SklearnCategoricalDensity(
LogisticRegression(max_iter=1000)
),
outcome_regressor=make_gps_outcome_pipeline(),
cv=2,
random_state=0,
)
.fit(X, t, y)
.predict(t=levels),
dtype=float,
).reshape(-1),
}
continuous_capable_benchmark = levels.with_columns(
*[
pl.Series(name, values)
for name, values in continuous_capable_results.items()
]
)
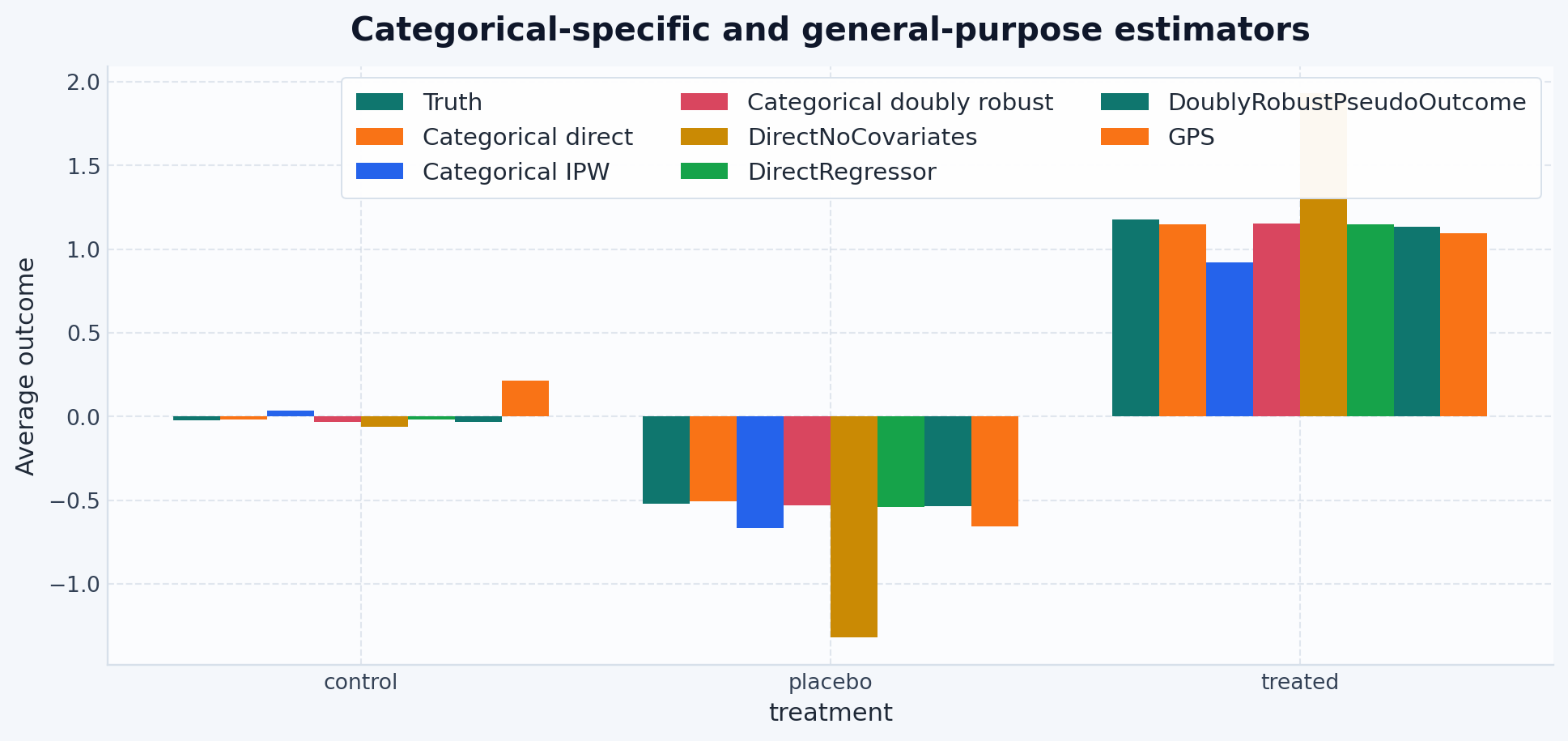
all_estimator_plot_results = {
"Truth": truth_curve,
"Categorical direct": direct_curve,
"Categorical IPW": ipw_curve,
"Categorical doubly robust": dr_curve,
"DirectNoCovariates": continuous_capable_results["DirectNoCovariates"],
"DirectRegressor": continuous_capable_results["DirectRegressor"],
"DoublyRobustPseudoOutcome": continuous_capable_results[
"DoublyRobustPseudoOutcome"
],
"GPS": continuous_capable_results["GPS"],
}
continuous_capable_benchmark